
BioSumm
Goals of the project
The BioSumm project tackles the problem of managing and exploiting the huge mass of information contained in increasingly wider text repositories such as PubMed. The project aims at becoming a powerful automatic instrument to support for both knowledge inference from scientific papers and biological validation of gene/proteins interactions obtained in different ways (e.g., with other data mining techniques).
Researchers that discover gene correlations by means of analysis tools (e.g., data mining tools) may exploit this framework to effectively support the biological validation of their results.
Framework description
BioSumm is a flexible and modular framework which analyzes large collections of unclassified biomedical texts and produces ad hoc summaries oriented to biological information. Its modular architecture is composed by two blocks:
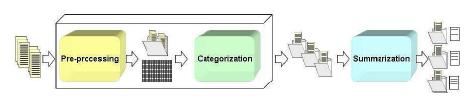
- Preprocessing and Categorization. It extracts relevant parts of the original document, produces a matricial representation of the sources and divides unclassified and rather diverse texts into homogeneous clusters.
- Summarization. For each cluster it produces a summary oriented to biological information.
The first block is a general purpose block with the goal of preparing the input documents for the summarization part. The prepocessing part is performed using the RapidMiner text plug-in whereas the categorization part exploits the CLUTO software package for clustering. The summarization block is the core of the framework. The summary generation is driven by a novel grading function, which biases sentence selection by means of an appropriate domain dictionary. In the current version of BioSumm, in order to focus on a biological target, the dictionary contains genes and proteins names and aliases.
Experimental results
BioSumm is neither a traditional summarizer nor a extractor of dictionary terms. It is designed to be a summarizer oriented to the biological domain. Thus, its summaries have both the expressive power of the traditional summaries and the domain specificity of documents produced by a dictionary entry extractor.
The difference with a traditional summarizer may be appreciated in the next table. It reports the six most graded sentences in BioSumm and in a traditional summary. The table was produced by the experiments carried on the scientific journals freely available in PubMed Central. Specifically, it contains sentences belonging to the a cluster of documents belonging to the Breast Cancer journal. The keywords of the cluster (the words describing its major topics) are proband, Ashkenazi, Jewish
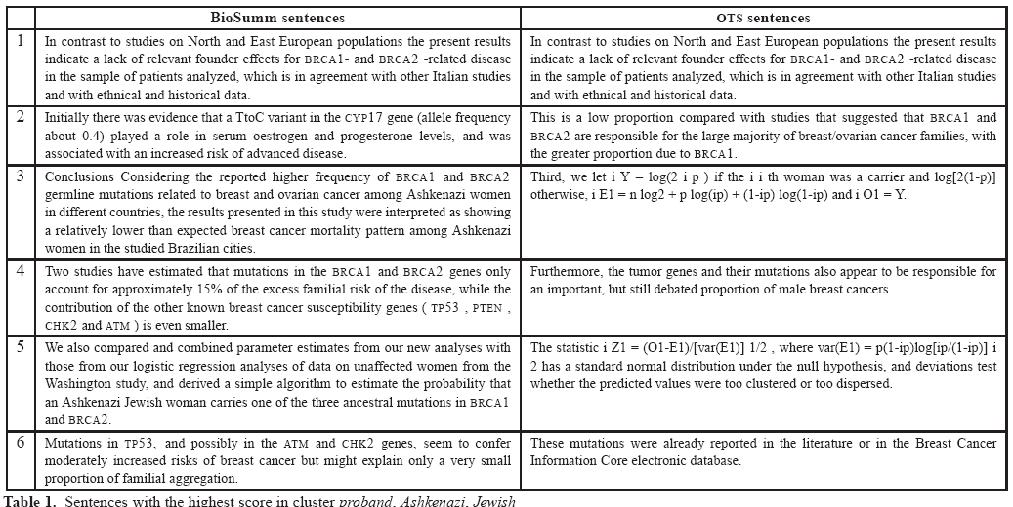
The comparison shows that BioSumm, although oriented on biology, is still able to cover all the major topics covered by a traditional summarizer. Moreover, its sentences are less generic and contains a lot of genes and proteins which are described in details and not only listed.
The results suggest that researchers that discover gene correlations by means of analysis tools (e.g., data mining tools) may exploit this framework to effectively support the biological validation of their results.
In the following some preliminary experimental results obtained by means of ROUGE are presented.
| Datasets | BioSumm | OTS | ||||
| ROUGE-2 | Precision | Recall | F-measure | Precision | Recall | F-measure |
| Breast Cancer | 0.08246 | 0.22553 | 0.11456 | 0.08026 | 0.21860 | 0.11141 |
| Arthritis Res | 0.09089 | 0.25362 | 0.12596 | 0.08844 | 0.24406 | 0.12197 |
| Datasets | BioSumm | OTS | ||||
| ROUGE-SU4 | Precision | Recall | F-measure | Precision | Recall | F-measure |
| Breast Cancer | 0.10038 | 0.28175 | 0.14053 | 0.09872 | 0.27599 | 0.13811 |
| Arthritis Res | 0.11095 | 0.31777 | 0.15498 | 0.10905 | 0.30888 | 0.15169 |
GUI Interface
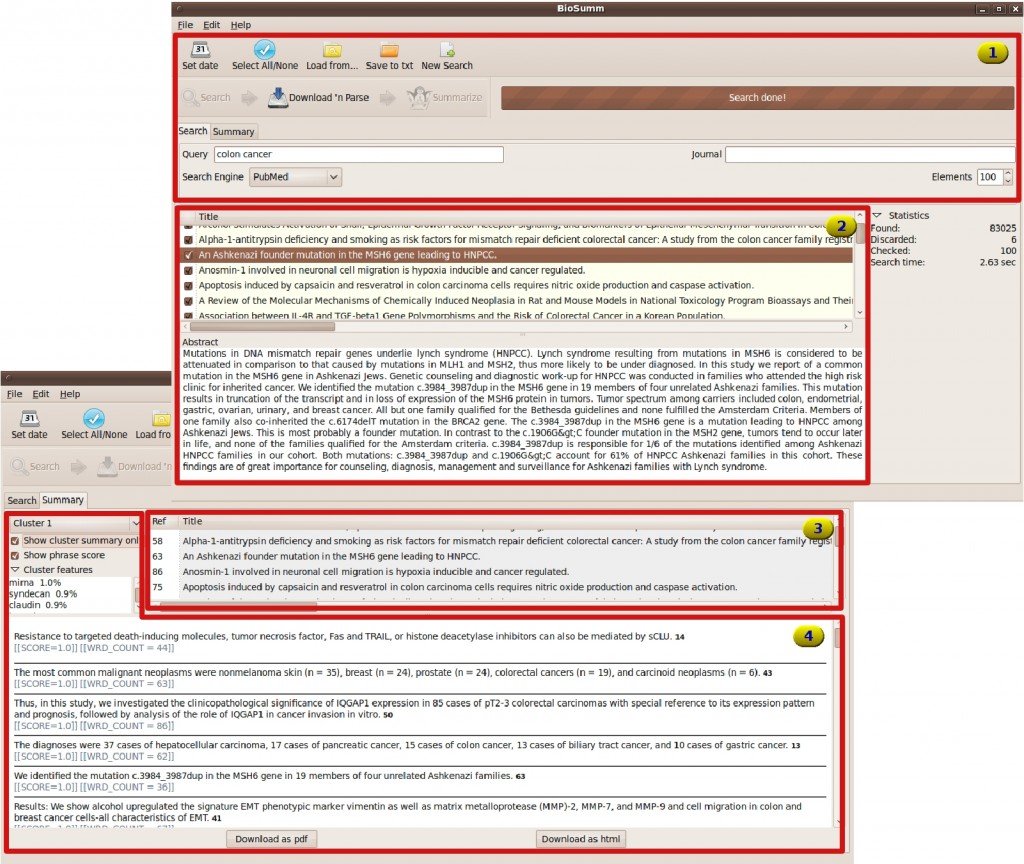
| Document search. The user can set the parameters to retrieve the documents from supported digital libraries. | |
| Document browsing. Management of retrieved documents to select the most relevant for summarization task. | |
| Documents of cluster. List of the documents belonging to a cluster identified by the clustering block. | |
| Cluster summary. The most relevant features (stems) which identify the topic of the cluster. |
Downloads
A new version of BioSumm will be avalaible
GUI application (32 bits) : BioSumm-1.1.tar
GUI application (64 bits) : BioSumm-1.1_64bit.tar
Presentation at Max Planck Institute (Saarbrucken)
Publications
For further details please refer to the scientific papers.
Contacts
Further details on the project may be obtained by contacting: